Data Set for Lung Imaging Challenge: Classification of findings in CT Lung Imaging
Download link: https://mosmed.ai/datasets/covid19_1110/
Click here to access the full document.
Dataset description:
- MosMedData includes 1,110 CT volumes collected from subjects in Moscow from 2020 March 1st to April 25th. This dataset contains deidentified human pulmonary CT scans with and without COVID-related radiological findings. At the beginning of the pandemic, CT served as a key tool to diagnose and monitor the progression of COVID-19 in Moscow. Clinical experts developed a procedure to grade the severity of COVID-19 based on CT radiological findings. The COVID-19 triage including follow-up by phone, admission to hospital or intensive care unit was decided by these severity findings along with other symptoms[1, 2].
- 1,110 subjects [age, (min, max, median), (18, 97, 47)] were recruited of which 42% were males, 56% females, 2% are other/unknown. They underwent a standard CT protocol using a Canon (Toshiba) Aquilion 64 CT scanner. The in-plane resolution is 0.8*0.8 mm2, the interslice distance is also 0.8 mm. However, this study only preserved every 10th slice of the original volume for storage. Therefore, the effective increment was 8mm[2]. The image matrix size is 512 x 512 x (36-41).
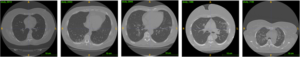
- Though there are several public datasets that have been used to investigate the application of deep learning in classifying COVID-19 findings in CT sliced images [3, 4], few studies focus on patient-wise, AI-based COVID-19 severity grading and categorical classification. The MosMedData provides 1,110 CT scans from non-repetitive subjects and corresponding 5-category annotations (ground truth). The grading of COVID-19 severity in CT was performed with a visual semi-quantitative scale adopted by the Russian Federation and used in Moscow hospitals. The dataset contains 254 scans without COVID-19 symptoms. The rest is split into 4 categories: CT1 (affected lung percentage 25% or below, 684 scans), CT2 (from 25% to 50%, 125 scans), CT3 (from 50% to 75%, 45 scans), CT4 (75% and above, 2 scans) (Figure 1). The final grade was decided based on the initial reading in clinics and a second reading by experts from the advisory department of the Center for Diagnostics and Telemedicine (CDT)[2].
- Dataset usage and previous studies:
We suggest utilizing the MosMedData to develop a volume-based deep learning model to identify the COVID-19 scans (binary classification) and evaluate the severity (categorical classification). Such an AI model is of great significance and of practical value for triage. - We suggest removing category CT4 as it only has 2 scans. Therefore, the final DL model will first identify the suspect COVID-19 CT scan and then classify the images into CT1 (mild), CT2 (moderate), and CT3 (severe). A patch/ROI/slice-based classification model plus a voting system may be sufficient. If memory allows, a model that directly takes the entire volume (matrix size is 512x512x38) as input can also be considered. Participants may refer to some memory-efficient networks.
- A previous study has explored using this dataset for binary classification (i.e., COVID vs. non-COVID), they achieved an AUC of 0.93[5], which implies that the categorical classification may be challenging but doable/feasible.
Figure 1. Examples of different COVID-19 severity. From left to right: CT0 (healthy), CT1 (mild), CT2 (moderate), CT3 (severe), CT4 (critical)
Performance Evaluation Criteria
1). The performance of each model will be judged by the number of correct classes (CT-0, CT-1, CT-2, CT-3) matched to the clinical/human expert classification. The results with the smallest classification error will be the winner.
2.) Results and programs of the model multi-classifier must be posted on GitHub to be considered in the competition.
3.) Each competitor/group should submit through the website, a 4-page paper summarizing the method and results, following the standard IEEE format for conference paper submissions. The report should provide the link to the GitHub post. The report should contain the proper citation/acknowledgment for the data use.
Registration
Register for this data challenge on the following page: https://easychair.org/conferences/?conf=ibacp2021
References
[1] S. Morozov et al., “Mosmeddata: Chest ct scans with covid-19 related findings dataset,” arXiv preprint arXiv:2005.06465, 2020.
[2] M. Goncharov et al., “CT-Based COVID-19 triage: Deep multitask learning improves joint identification and severity quantification,” Medical Image Analysis, vol. 71, p. 102054, 2021/07/01/ 2021, doi: https://doi.org/10.1016/j.media.2021.102054.
[3] E. Soares, P. Angelov, S. Biaso, M. H. Froes, and D. K. Abe, “SARS-CoV-2 CT-scan dataset: A large dataset of real patients CT scans for SARS-CoV-2 identification,” medRxiv, p. 2020.04.24.20078584, 2020, doi: 10.1101/2020.04.24.20078584.
[4] X. Yang, X. He, J. Zhao, Y. Zhang, S. Zhang, and P. Xie, “COVID-CT-dataset: a CT scan dataset about COVID-19,” arXiv preprint arXiv:2003.13865, 2020.
[5] C. Jin et al., “Development and evaluation of an artificial intelligence system for COVID-19 diagnosis,” Nature Communications, vol. 11, no. 1, p. 5088, 2020/10/09 2020, doi: 10.1038/s41467-020-18685-1.
Relevant Articles:
Development and evaluation of an artificial intelligence system for COVID-19 diagnosis
MOSMEDDATA_CHEST CT SCANS WITH COVID-19 RELATED FINDINGS DATASET